See full list on marketplace.visualstudio.com. Feb 10, 2021 Markdown is a markup language commonly used to simplify the process of writing content in an easy-to-read text format, which a software tool or programming library can convert into HTML to display in a browser or another writing program. With Jekyll, you’ll typically work with content in Markdown, a lightweight markup language designed for text formatting. The Liquid templating engine is used to place this Markdown content into a HTML template, and to combine templates representing various parts of a page (say, header, footer and content) in a modular and re-usable manner. Showdown is a Javascript Markdown to HTML bidirectional converter, based on the original works by John Gruber. Showdown can be used client side (in the browser) or server side (with nodejs). See it working in the live demo! Var converter = new showdown.Converter. Free online markdown to HTML converter. Markdown CheatSheet # 1. Basic # H1 ## H2 ### H3 #### H4 ##### H5 ##### H6 H1 H2 -.italics. or italics.
In this tutorial, we will cover the R
knitr package that is used to convertR Markdown into a rendered document (HTML, PDF, etc).Learning Objectives
At the end of this activity, you will:
- Be able to produce (‘knit’) an HTML file from a R Markdown file.
- Know how to modify chunk options to change the output in your HTML file.
Things You’ll Need To Complete This Tutorial
You will need the most current version of R and, preferably, RStudio loaded onyour computer to complete this tutorial.
Install R Packages
- knitr:
install.packages('knitr') - rmarkdown:
install.packages('rmarkdown')
Share & Publish Results Directly from Your Code!
The
knitr package allow us to:- Publish & share preliminary results with collaborators.
- Create professional reports that document our workflow and results directlyfrom our code, reducing the risk of accidental copy and paste or transcription errors.
- Document our workflow to facilitate reproducibility.
- Efficiently change code outputs (figures, files) given changes in the data, methods, etc.
Publish from Rmd files with knitr
To complete this tutorial you need:
- The R
knitrpackage to complete this tutorial. If you need help installing packages, visit the R packages tutorial. - An R Markdown document that contains a YAML header, code chunks and markdownsegments. If you don't have an .Rmd file, visit the R Markdown tutorial to create one.
When To Knit: Knitting is a useful exercisethroughout your scientific workflow. It allows you to see what your outputslook like and also to test that your code runs without errors.The time required to knit depends on the length and complexity of the scriptand the size of your data.
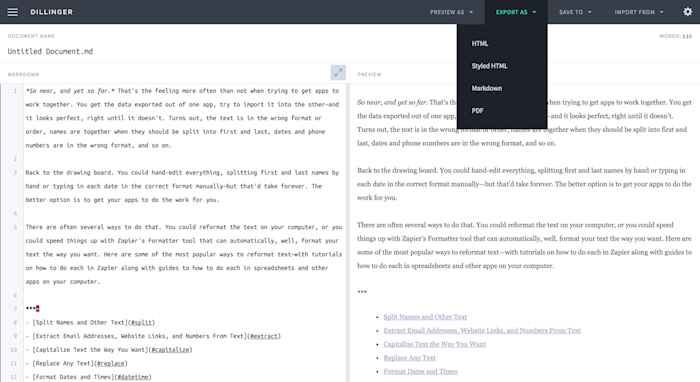
How to Knit
To knit in RStudio, click the knit pull down button. You want to use the
knit HTML for this lesson.
knit HTML for this lesson.
Wavepad audio editor full. When you click the Knit HTML button, a window will open in your console titled R Markdown. Thispane shows the knitting progress. The output (HTML in this case) file willautomatically be saved in the current working directory. If there is an errorin the code, an error message will appear with a line number in the R Consoleto help you diagnose the problem.
Data Tip: You can run
knitr from the command promptusing: render(“input.Rmd”, “all”).Activity: Knit Script
Knit the .Rmd file that you built inthe last tutorial.What does it look like?
View the Output
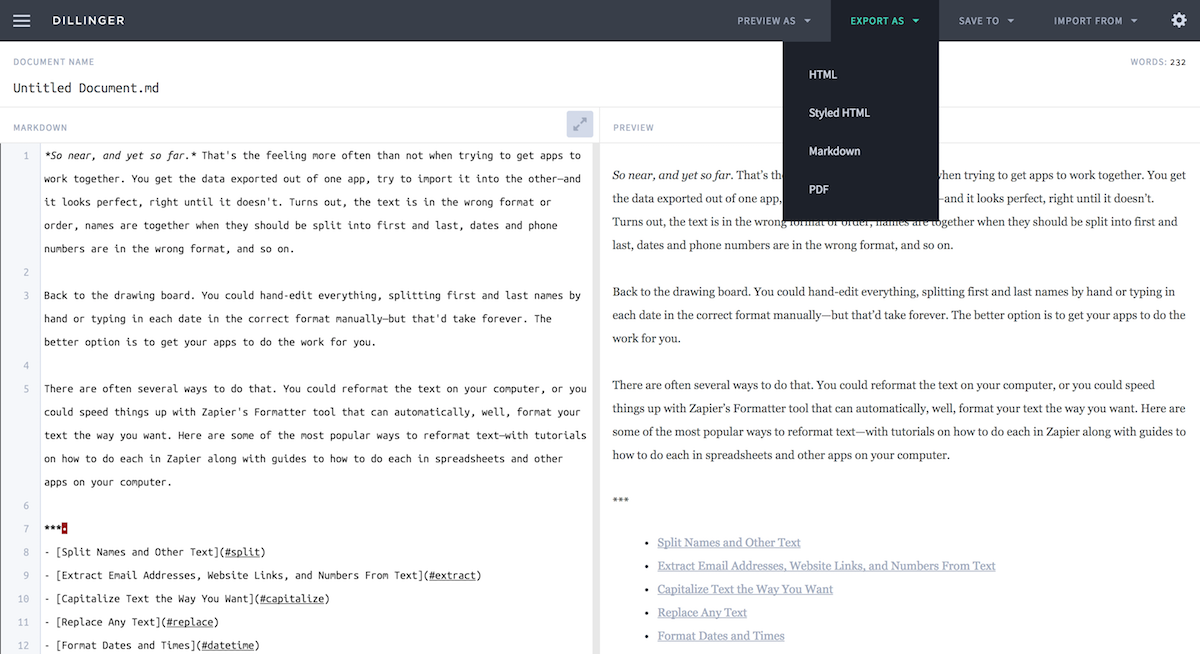
When knitting is complete, the new HTML file produced will automatically open.
Notice that information from the YAML header (title, author, date) is printedat the top of the HTML document. Then the HTML shows the text, code, andresults of the code that you included in the RMD document.
Markdown To Html Windows
Data Institute Participants: Complete Week 2 Assignment
Convert Word To Markdown
- Read this week’s assignment closely.
- Be sure to carefully check your knitr output to make sure it is rendering theway you think it should!
- When you are complete, submit your .Rmd and .html files to the NEON Institute participants GitHub repository (NEONScience/DI-NEON-participants).
- The files will have automatically saved to your R working directory, you will need to transfer the files to the /participants/pre-institute3-rmd/ directory and submitted via a pull request.